Marine Biology

Boston University has a world-class program in marine biology that is active in training students at both the undergraduate and graduate level. The Marine Biology research group includes professors who are leaders in their subdisciplines, including evolutionary and conservation genetics of marine organisms, sensory biology, ichthyology, evolution and development of marine organisms, marine microbial ecology, marine community ecology, and marine conservation science.
Marine Biology faculty and their students are actively engaged in such exciting research topics as biomimicry, functional genomics, ocean exploration, and marine conservation, and are working at the cutting edge of scientific efforts to combat overfishing, global climate change, and marine species extinction. Our research is strengthened by close ties between Biology faculty and colleagues from the Department of Earth & Environment . In addition, both faculty and students enjoy the benefits of Boston University’s formal partnerships with the New England Aquarium , the Stellwagen Bank National Marine Sanctuary , the Sea Education Association , and a variety of government and non-governmental marine conservation organizations.

Faculty with Related Research

Peter Buston , Associate Professor of Biology; Director of the BU Marine Program
Evolutionary Ecology, Animal Behavior, Marine Ecology and Biological Oceanography.

Sarah W. Davies , Associate Professor of Biology
ecological genomics, population genetics, molecular ecology, coral ecology, climate change, symbiosis
Ethan Deyle , Research Assistant Professor of Biology
Quantitative Ecology, Environmental Data Science, Nonlinear Dynamics, Applied Complex Systems, Marine Ecology

John R. Finnerty , Associate Professor of Biology
Evolutionary and ecological developmental biology; evolutionary and ecological genomics; marine biodiversity; global change biology; coral conservation.

Robinson W. Fulweiler , Professor of Biology (jointly with Earth & Environment)
Biogeochemistry and Marine Ecology
Thomas Gilmore , Professor of Biology
molecular biology, cell biology, signal transduction, cancer, molecular ecology

Les Kaufman , Professor of Biology
marine biology, evolutionary ecology, and conservation biology

Phillip S. Lobel , Professor of Biology
ichthyology; behavioral ecology and taxonomy of fishes

Ranjan Muthukrishnan , Research Assistant Professor
community ecology, marine ecology, coral reefs, invasive species, ecological modeling
EBE and Marine Biology

Randi Rotjan , Research Associate Professor of Biology; Senior Lecturer; Director of Masters Studies
Marine ecology, conservation biology, behavioral ecology, organismal physiology, coral reefs
2025 Advanced Research Training Course Applications Are Now Open!

The Marine Biological Laboratory is a world-renowned scientific institution, and scientists and students come to the MBL throughout the year to pursue research and education in diverse areas of fundamental biological discovery.
The Division of Research encompasses all research programs at the Laboratory, including the development and management of research services, research budgets, the assignment of research space, the disbursement of research awards and fellowships, training and education for the responsible conduct of scientific research, IACUC, IRB, and guidance for graduate and postdoctoral trainees.
The Division also works closely with the Office of the President/Director to develop new strategic research initiatives and to manage all faculty appointments, promotions, and mentoring.
Our Research Centers
Research at the MBL takes place throughout the year in a number of research centers:
Josephine Bay Paul Center for Comparative Molecular Biology & Evolution
Eugene Bell Center for Regenerative Biology and Tissue Engineering
The Ecosystems Center
Whitman Center

Research resources to support ongoing discovery at the MBL.
Marine Resources Center
National Xenopus Resource (NXR)
The Microbiome Center
Imaging Resources
Central Microscopy Facility
The newly established joint program between the Marine Biological Laboratory (MBL) and the University of Chicago (UChicago) leverages the unique partnership between two leading research institutions and combines the best of both worlds – access to a collaborative and expansive research environment that spans the scales of biological discovery at the MBL in Woods Hole, MA and the first-class resources of the University of Chicago.
MBL has several funding opportunities for those who wish to come to the MBL to do research either collaboratively or individually throughout the year:
Whitman Center Fellowships
Edwin Barbey Charitable Trust Fellowship
Grass Foundation Fellowships
Faculty and Whitman Scientists
Researchers and Support Staff
MBL Fellows
MBL Postdoctoral Association
Everything you need to get started at the MBL.
Campus Information
Come Back to the MBL
Animal Care and Support
Equipment, Supplies and Facilities
Research Policies and Procedures
Order An Organism
Research Administration
Office of Sponsored Programs
Health and Safety
MBLWHOI Library
Scientific Publications
Captured by MBL Hibbitt Fellow Caroline Albertin, this video shows California two-spot octopuses (Octopus bimaculoides) emerging from their egg casings.
Thank you for visiting nature.com. You are using a browser version with limited support for CSS. To obtain the best experience, we recommend you use a more up to date browser (or turn off compatibility mode in Internet Explorer). In the meantime, to ensure continued support, we are displaying the site without styles and JavaScript.
- View all journals
- Explore content
- About the journal
- Publish with us
- Sign up for alerts
Marine biology articles within Scientific Reports
Article 11 November 2024 | Open Access
Enhancing reef carbonate budgets through coral restoration
- Emily Esplandiu
- , John Morris
- & Diego Lirman
Article 03 November 2024 | Open Access
Desert dust improves the photophysiology of heat-stressed corals beyond iron
- Katherine Amorim
- , R. Grover
- & Christine Ferrier-Pagès
Article 02 November 2024 | Open Access
Development of a method for estimating asari clam distribution by combining three-dimensional acoustic coring system and deep neural network
- Tokimu Kadoi
- , Katsunori Mizuno
- & Kei Terayama
Investigating the potential effect of Holothuria scabra extract on osteogenic differentiation in preosteoblast MC3T3-E1 cells
- Sineenart Songkoomkrong
- , Siriporn Nonkhwao
- & Prateep Amonruttanapun
Article 01 November 2024 | Open Access
Noninvasive, epigenetic age estimation in an elasmobranch, the cownose ray ( Rhinoptera bonasus )
- D. Nick Weber
- , Jennifer T. Wyffels
- & David S. Portnoy
Article 29 October 2024 | Open Access
Seaweed functional strategies, functional groups, and taxon dynamics through a 213-year historical series of Rio De Janeiro Bay
- João P. G. Machado
- & Vinícius P. Oliveira
Article 28 October 2024 | Open Access
Population changes in a Southern Ocean krill predator point towards regional Antarctic sea ice declines
- Matthew Germishuizen
- , Marcello Vichi
- & Els Vermeulen
Article 22 October 2024 | Open Access
Seabird nutrients increase coral calcification rates and boost reef carbonate production
- Ines D. Lange
- & Cassandra E. Benkwitt
Acute copper stress showed toxic effects on the physiological metabolism of Ulva lactuca , a common green macroalgae
- Weimin Chen
- & Manying Sun
Article 17 October 2024 | Open Access
Hydromechanical properties of metachronal swimming in polychaetes
- Brad J. Gemmell
- , Sean P. Colin
- & John H. Costello
Article 16 October 2024 | Open Access
Bacterial and Symbiodiniaceae communities’ variation in corals with distinct traits and geographical distribution
- Livia Bonetti Villela
- , Arthur Weiss da Silva-Lima
- & Paulo Sergio Salomon
Article 12 October 2024 | Open Access
Vertebrate endocrine disruptors induce sex-reversal in blue mussels
- K. Garrett Evensen
- , Emily Rusin
- & Helen C. Poynton
Article 10 October 2024 | Open Access
Factors shaping pelagic-benthic coupling in the process of settlement in an Arctic fjord
- Marta Ronowicz
- , Piotr Balazy
- & Agata Weydmann-Zwolicka
Article 09 October 2024 | Open Access
Acoustic radiation characteristics of shark skin inspired surface modified plates
- & Ritwik Ghoshal
Integrated physiological response by four species of Rhodophyta to submarine groundwater discharge reveals complex patterns among closely-related species
- Veronica L. Gibson
- , Angelene Dedloff
- & Celia M. Smith
Article 06 October 2024 | Open Access
Sex-specific vertical movements of spawning atlantic cod in coastal habitats inferred from acoustic telemetry
- J. E. Skjæraasen
- , E. M. Olsen
- & L. D. Sivle
Article 03 October 2024 | Open Access
Physalia gonodendra are not yet sexually mature when released
- Kohei Oguchi
- , Gaku Yamamoto
- & Casey W. Dunn
Article 28 September 2024 | Open Access
Endemic, cosmopolitan, and generalist taxa and their habitat affinities within a coastal marine microbiome
- Chase C. James
- , Andrew E. Allen
- & Andrew D. Barton
Population genomic structure of the sea urchin Diadema africanum , a relevant species in the rocky reef systems across the Macaronesian archipelagos
- Marc Peralta-Serrano
- , José Carlos Hernández
- & Rocio Pérez-Portela
The distribution of seaweed forms and foundational assumptions in seaweed biology
Article 27 September 2024 | Open Access
Establishing complexity targets to enhance artificial reef designs
- Elisabeth Riera
- , Benjamin Mauroy
- & Cédric Hubas
Determination of biogeochemical properties in sea waters using the inversion of the three-stream irradiance model
- Paolo Lazzari
- , Mirna Gharbi Dit Kacem
- & Vincenzo Vellucci
Article 20 September 2024 | Open Access
Oceanographical-driven dispersal and environmental variation explain genetic structure in an upwelling coastal ecosystem
- Lívia Peluso
- , Juan Faúndez
- & Pablo Saenz-Agudelo
Article 18 September 2024 | Open Access
A portable multi-taxa phenotyping device to retrieve physiological performance traits
- Hadley England
- , Andrei Herdean
- & Emma F. Camp
Article 17 September 2024 | Open Access
Diverse baleen whale acoustic occurrence around two sub-Antarctic islands: A tale of residents and visitors
- Fannie W. Shabangu
- , Tessa Munoz
- & Tarron Lamont
Fisheries shocks provide an opportunity to reveal multiple recruitment sources of sardine in the Sea of Japan
- Tatsuya Sakamoto
- , Motomitsu Takahashi
- & Toyoho Ishimura
Article 15 September 2024 | Open Access
Availability to predators and a size structure of the Antarctic krill Euphausia superba in the 48.1 CCAMLR subarea
- Anna Panasiuk
- , Gabriela Gic-Grusza
- & Małgorzata Korczak-Abshire
Article 13 September 2024 | Open Access
Physiological resilience of intertidal chitons in a persistent upwelling coastal region
- Carolina Fernández
- , María Josefina Poupin
- & Marco Antonio Lardies
Article 08 September 2024 | Open Access
Leveraging deep learning and computer vision technologies to enhance management of coastal fisheries in the Pacific region
- George Shedrawi
- , Franck Magron
- & Neil L. Andrew
Article 06 September 2024 | Open Access
From ctenophores to scyphozoans: parasitic spillover of a burrowing sea anemone
- Anastasiia Iakovleva
- , Arseniy R. Morov
- & Tamar Guy-Haim
Article 04 September 2024 | Open Access
Predicting 3D and 2D surface area of corals from simple field measurements
- Josie F. Chandler
- , Will F. Figueira
- & Morgan S. Pratchett
Article 31 August 2024 | Open Access
Struggling with fish age, a comparison of otolith preparation techniques to unravel age and growth of boarfish, Capros aper (Linnaeus, 1758)
- Maria Inês Silva
- , Rui Martins
- & Ana Rita Vieira
Article 29 August 2024 | Open Access
Sex-specific differences in the growth and population characteristics of Sand crab Ovalipes punctatus (De Haan, 1833) in coastal waters of Korea
- Hyeon Gyu Lee
- , Jae Mook Jeong
- & Youn Hee Choi
Article 28 August 2024 | Open Access
Customized digestion protocols for copepods, euphausiids, chaetognaths and fish larvae facilitate the isolation of ingested microplastics
- Imke Podbielski
- , Thea Hamm
- & Mark Lenz
Article 26 August 2024 | Open Access
Insights into aging mechanisms from comparative genomics in orange and silver roughies
- Dido Carrero
- , Maria Pascual-Torner
- & Carlos López-Otín
Multi-marker DNA metabarcoding for precise species identification in ichthyoplankton samples
- André O. Ferreira
- , Olga M. Azevedo
- & Filipe O. Costa
Article 20 August 2024 | Open Access
Habitat for Coilia nasus in southern Zhejiang Province, China, based on a maximum entropy model
- & Weicheng Liu
Article 19 August 2024 | Open Access
Genetics and ontogeny are key factors influencing thermal resilience in a culturally and economically important bivalve
- Natalí J. Delorme
- , Nick King
- & Norman L. C. Ragg
Article 11 August 2024 | Open Access

Epiphytic microbiome associated with intertidal seaweeds in the Mediterranean Sea: comparative analysis of bacterial communities across seaweed phyla
- , Álvaro Israel
- & Tal Luzzatto-Knaan
Article 10 August 2024 | Open Access
Functional prediction based on 16S rRNA metagenome data from bacterial microbiota associated with macroalgae from the Peruvian coast
- Bianca E. Vigil
- , Francisco Ascue
- & Danilo E. Bustamante
Article 03 August 2024 | Open Access
Sterolysin from a 1950s culture of Karlodinium veneficum (aka Gymnodinium veneficum Ballantine) forms lethal sterol dependent membrane pores
- Allen R. Place
- , Josefina Ramos-Franco
- & Mark T. Hamann
Article 30 July 2024 | Open Access
Response of hypoxia to future climate change is sensitive to methodological assumptions
- Kyle E. Hinson
- , Marjorie A. M. Friedrichs
- & Hanqin Tian
Article 29 July 2024 | Open Access
Mark-recapture validates the use of photo-identification for the widely distributed blue-spotted ribbontail ray, Taeniura lymma
- Ashlie J. McIvor
- , Collin T. Williams
- & Michael L. Berumen
Article 20 July 2024 | Open Access
Vertical and horizontal environmental DNA (eDNA) patterns of fish in a shallow and well-mixed North Sea area
- Nergiz Dukan
- , Isolde Cornelis
- & Sofie Derycke
Article 15 July 2024 | Open Access
Personal electric deterrents can reduce shark bites from the three species responsible for the most fatal interactions
- Thomas M. Clarke
- , Adam Barnett
- & Charlie Huveneers
Article 02 July 2024 | Open Access
Effects of microplastics on reproductive characteristics and mechanisms of the marine rotifer Brachionus plicatilis
- Taekyoung Seong
- , Sae Yamamoto
- & Hee-Jin Kim
Article 01 July 2024 | Open Access
Ocean warming and acidification adjust inter- and intra-specific variability in the functional trait expression of polar invertebrates
- Thomas J. Williams
- , Adam J. Reed
- & Martin Solan
Article 27 June 2024 | Open Access
Marine heatwaves alter the nursery function of coastal habitats for juvenile Gulf of Alaska Pacific cod
- Hillary L. Thalmann
- , Benjamin J. Laurel
- & Jessica A. Miller
Article 24 June 2024 | Open Access
The absence of canonical respiratory complex I subunits in male-type mitogenomes of three Donax species
- Artur Burzyński
- , Beata Śmietanka
- & Marek Lubośny
Article 17 June 2024 | Open Access
Spatial prioritization of dugong habitats in India can contribute towards achieving the 30 × 30 global biodiversity target
- , Sharad Bayyana
- & Jeyaraj Antony Johnson
Browse broader subjects
- Ocean sciences
Quick links
- Explore articles by subject
- Guide to authors
- Editorial policies
Articles on Marine biology
Displaying 1 - 20 of 180 articles.

As the oceans warm, deep-living algae are thriving – with major potential effects for the marine ecosystem
Johan Viljoen , University of Exeter ; Bob Brewin , University of Exeter , and Xuerong Sun , University of Exeter

Sharks and rays leap out of the water for many reasons, including feeding, courtship and communication
A. Peter Klimley , University of California, Davis

Whales are recovering from near extinction, but industrial fishing around Antarctica competes for their sole food source
Matthew Savoca , Stanford University

Squid have tiny teeth in their suckers − scientists could use their unique properties to make self-healing materials
Abdon Pena-Francesch , University of Michigan

Sea lions wearing cameras and trackers map new habitats
Nathan Angelakis , University of Adelaide

Oceans without sharks would be far less healthy – new research
Michael Heithaus , Florida International University

From glowing corals to vomiting shrimp, animals have used bioluminescence to communicate for millions of years – here’s what scientists still don’t know about it
Danielle DeLeo , Florida International University and Andrea Quattrini , Smithsonian Institution

Climate change is causing marine ‘coldwaves’ too, killing wildlife
Nicolas Benjamin Lubitz , James Cook University and David Schoeman , University of the Sunshine Coast

Sex, birth and whalesong: life on the humpback highway
Vanessa Pirotta , Macquarie University

As climate change and pollution imperil coral reefs, scientists are deep-freezing corals to repopulate future oceans
Mary Hagedorn , Smithsonian Institution

How do halibut migrate? Clues are in their ear bones
Charlotte Gauthier , Université du Québec à Chicoutimi (UQAC)

Surviving fishing gear entanglement isn’t enough for endangered right whales – females still don’t breed afterward
Joshua Reed , Macquarie University ; Leslie New , Ursinus College ; Peter Corkeron , Griffith University , and Rob Harcourt , Macquarie University
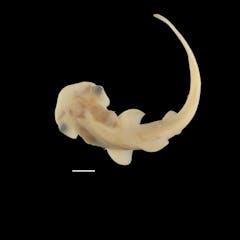
Rare access to hammerhead shark embryos reveals secrets of its unique head development
Gareth J. Fraser , University of Florida

‘Jaws’ portrayed sharks as monsters 50 years ago, but it also inspired a generation of shark scientists
Gavin Naylor , University of Florida

Not all underwater reefs are made of coral − the US has created artificial reefs from sunken ships, radio towers, boxcars and even voting machines
Avery Paxton , National Oceanic and Atmospheric Administration and D'amy Steward , University of Guam

I set out to investigate where silky sharks travel − and by chance documented a shark’s amazing power to regenerate its sabotaged fin
Chelsea Black , University of Miami

What happens to the ocean if we take out all the fish? A marine ecologist explains the complex roles fish play in their ecosystem
Kory Evans , Rice University

Shipwrecks teem with underwater life, from microbes to sharks
Avery Paxton , National Oceanic and Atmospheric Administration

As seas get warmer, tropical species are moving further from the equator
Karolina Zarzyczny , University of Southampton

Australian dolphins have the world’s highest concentrations of ‘forever chemicals’
Chantel Foord , RMIT University
Related Topics
- Biodiversity
- Climate change
- Conservation
- Coral reefs
- Marine conservation
- Marine ecosystems
- Marine life
Top contributors
Professor, Macquarie University
Executive Dean of the College of Arts, Sciences & Education and Professor of Biological Sciences, Florida International University
Senior Curator of Marine Invertebrates, Museums Victoria Research Institute
Professor of Marine Botany, The University of Western Australia
Faculty Research Associate in Marine Biology, Arizona State University
Royal Society University Research Fellow, University of Sheffield
Founding Professor of the Minderoo-UWA Deep-Sea Research centre, The University of Western Australia
Postdoctoral associate, University of Florida
Assistant Professor, University of Rhode Island
Professor, Marine Biology, University of Adelaide
Lecturer and Researcher in Behavioural ecology and Cephalopod Biology, Anglia Ruskin University
Director, Florida Program for Shark Research, University of Florida
PhD Researcher in Marine Ecology, University of Exeter
Professor of Biology and Director, Kewalo Marine Laboratory, University of Hawaii
Professor in marine ecology, UNSW Sydney
- X (Twitter)
- Unfollow topic Follow topic
Marine Biology Research

Subject Area and Category
- Aquatic Science
- Ecology, Evolution, Behavior and Systematics
- Oceanography
Taylor and Francis Ltd.
Publication type
17451000, 17451019
Information
How to publish in this journal
The set of journals have been ranked according to their SJR and divided into four equal groups, four quartiles. Q1 (green) comprises the quarter of the journals with the highest values, Q2 (yellow) the second highest values, Q3 (orange) the third highest values and Q4 (red) the lowest values.
The SJR is a size-independent prestige indicator that ranks journals by their 'average prestige per article'. It is based on the idea that 'all citations are not created equal'. SJR is a measure of scientific influence of journals that accounts for both the number of citations received by a journal and the importance or prestige of the journals where such citations come from It measures the scientific influence of the average article in a journal, it expresses how central to the global scientific discussion an average article of the journal is.
Evolution of the number of published documents. All types of documents are considered, including citable and non citable documents.
This indicator counts the number of citations received by documents from a journal and divides them by the total number of documents published in that journal. The chart shows the evolution of the average number of times documents published in a journal in the past two, three and four years have been cited in the current year. The two years line is equivalent to journal impact factor ™ (Thomson Reuters) metric.
Evolution of the total number of citations and journal's self-citations received by a journal's published documents during the three previous years. Journal Self-citation is defined as the number of citation from a journal citing article to articles published by the same journal.
Evolution of the number of total citation per document and external citation per document (i.e. journal self-citations removed) received by a journal's published documents during the three previous years. External citations are calculated by subtracting the number of self-citations from the total number of citations received by the journal’s documents.
International Collaboration accounts for the articles that have been produced by researchers from several countries. The chart shows the ratio of a journal's documents signed by researchers from more than one country; that is including more than one country address.
Not every article in a journal is considered primary research and therefore "citable", this chart shows the ratio of a journal's articles including substantial research (research articles, conference papers and reviews) in three year windows vs. those documents other than research articles, reviews and conference papers.
Ratio of a journal's items, grouped in three years windows, that have been cited at least once vs. those not cited during the following year.
Evolution of the percentage of female authors.
Evolution of the number of documents cited by public policy documents according to Overton database.
Evolution of the number of documents related to Sustainable Development Goals defined by United Nations. Available from 2018 onwards.
Leave a comment
Name * Required
Email (will not be published) * Required
* Required Cancel
The users of Scimago Journal & Country Rank have the possibility to dialogue through comments linked to a specific journal. The purpose is to have a forum in which general doubts about the processes of publication in the journal, experiences and other issues derived from the publication of papers are resolved. For topics on particular articles, maintain the dialogue through the usual channels with your editor.

Follow us on @ScimagoJR Scimago Lab , Copyright 2007-2024. Data Source: Scopus®

Cookie settings
Cookie Policy
Legal Notice
Privacy Policy

IMAGES
VIDEO
COMMENTS
In its first ten years, MBRJ has published nearly 700 submissions from 55 countries, including many research and review articles as well as eight Thematic Issues. Marine Biology Research covers a broad range of topics, including: Ecology. Autotrophs and Heterotrophs, Food webs. Behaviour.
Marine biology is the study of life in the oceans and brackish waters, which ranges from archaea and bacteria to marine mammals, and includes organisms such as corals that affect the shape of...
Explore the current issue of Marine Biology Research, Volume 20, Issue 5-6, 2024.
Ocean sciences span the physics, chemistry, and biology of marine systems. The field encompasses ocean circulation, energy dissipation, marine biology, ecology, biogeochemical cycles, water...
fundamental biological research, often using marine organisms as novel model systems, encompassing research in regenerative biology, neuroscience, sensory physiology, and comparative evolution and genomics; the study of microbiomes and microbial diversity and ecology in a variety of ocean and terrestrial habitats; imaging and computation;
The Marine Biology research group includes professors who are leaders in their subdisciplines, including evolutionary and conservation genetics of marine organisms, sensory biology, ichthyology, evolution and development of marine organisms, marine microbial ecology, marine community ecology, and marine conservation science.
The Marine Biological Laboratory is a world-renowned scientific institution, and scientists and students come to the MBL throughout the year to pursue research and education in diverse areas of fundamental biological discovery.
Read the latest Research articles in Marine biology from Scientific Reports.
From glowing corals to vomiting shrimp, animals have used bioluminescence to communicate for millions of years – here’s what scientists still don’t know about it. Danielle DeLeo, Florida...
Marine Biology Research ( MBRJ) provides a worldwide forum for key information, ideas and discussion on all areas of marine biology and biological oceanography. Founded in 2005 as a merger of two Scandinavian journals, Sarsia and Ophelia, MBRJ is based today at the Institute of Marine Research, Bergen, Norway.